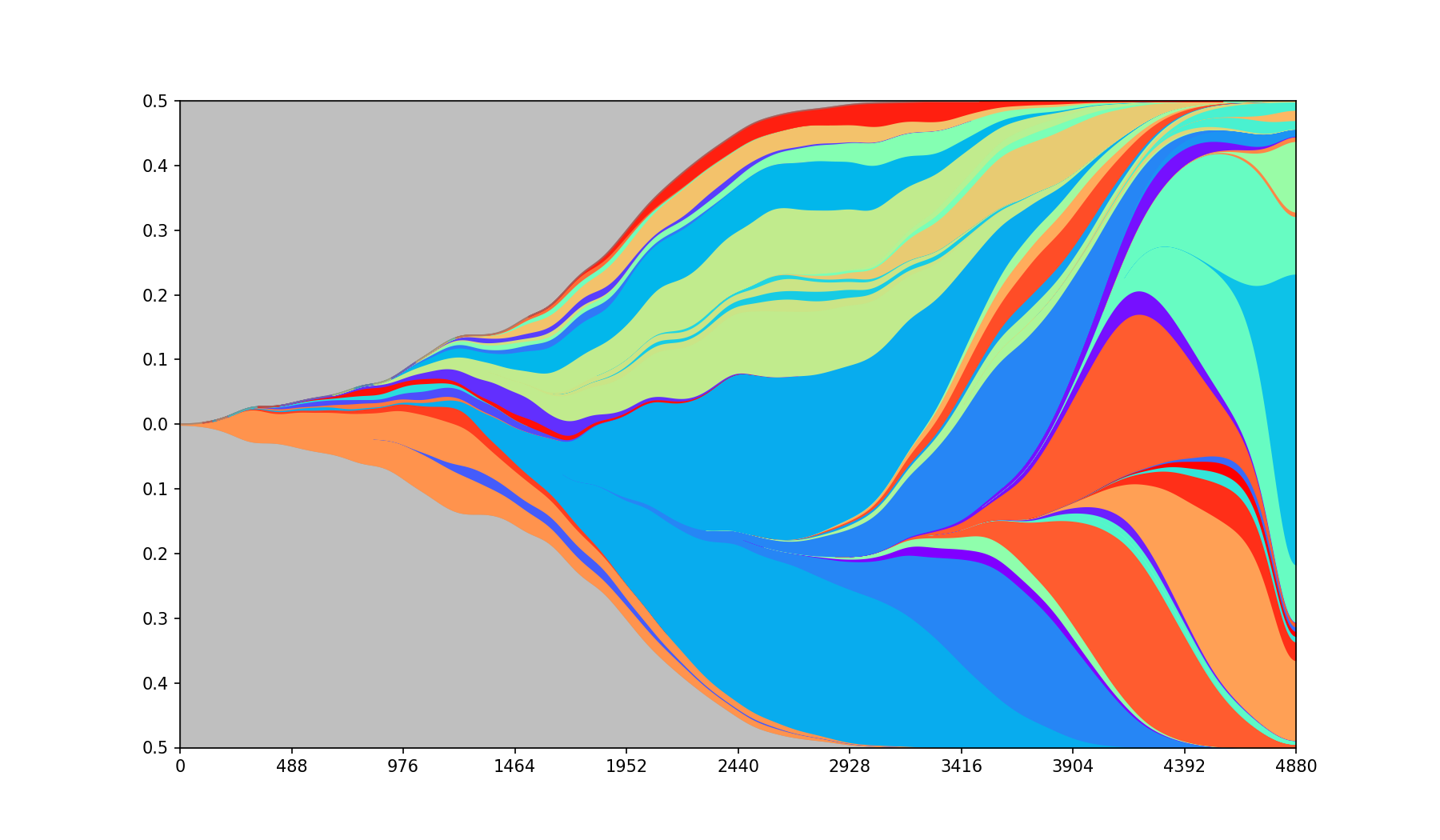
PyFish is a Python 3 package for creation of Fish (Muller) plots like the one below.
- polynomial interpolation
- curve smoothing
- high performance
- works with low and high density data
PyFish can be used either as a stand-alone tool or as a plotting library.
PyFish requires Python >= 3.8
The package can be installed using Conda (from the bioconda channel)
conda install -c bioconda pyfish
or Pip
pip install pyfish.
The program takes two tables:
- one describing the size of individual subgroups at given points in time, referred to as populations,
- one describing the parent-child relationships between the subgroups, referred to as parent tree.
Populations table has the schema (Id: +int, Step: +int, Pop: +int), where:
Idis a numerical identifier of a subgroup,Stepis a natural ordinal describing the logical time when the population is measured,Popis the size of the population of the subgroup at the given step.
An example populations table:
| Id | Step | Pop |
|---|---|---|
| 0 | 0 | 100 |
| 0 | 1 | 40 |
| 0 | 2 | 20 |
| 0 | 3 | 0 |
| 1 | 0 | 10 |
| 1 | 3 | 50 |
| 1 | 5 | 100 |
| 2 | 4 | 20 |
| 2 | 5 | 50 |
| 3 | 0 | 10 |
| 3 | 1 | 20 |
| 3 | 5 | 10 |
Parent tree has the schema (ParentId: +int, ChildId: +int), where:
ParentIdis an id matching the population table,ChildIdis an id matching the population table describing the direct progeny of the parent.
An example parent tree:
| ParentId | ChildId |
|---|---|
| 0 | 1 |
| 1 | 2 |
| 0 | 3 |
Note: If multiple nodes have no parent, or if population IDs are not listed in the parent tree, a synthetic root with zero population is automatically created to parent them all.
A synthetic root is always centered and has a width zero, even when using curved or separate modes.
We provide example data. From the root folder of the project call:
pyfish tests/populations.csv tests/parent_tree.csv out.png
This will create a plot called out.png in the folder.
Additional execution parameters are described below.
The populations and parent_tree tables can be constructed directly as dataframes.
A worked API example covering common options (raw, curved, separated, interpolated, and colormap comparison) is provided in example.ipynb.
The library contains three public functions:
process_dataTakes the input data and parameters and creates data suitable for plotting. Additional arguments match the parameters as described below.setup_figureResizes the figure and adds labels for axes.fish_plotCalls the plotting function on the input parameters.
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from pyfish import fish_plot, process_data, setup_figure
populations = np.array([[0, 0, 100], [0, 1, 40], [0, 2, 20], [0, 3, 0], [1, 0, 10], [1, 3, 50],
[1, 5, 100], [2, 4, 20], [2, 5, 50], [3, 0, 10], [3, 1, 20], [3, 5, 10]])
parent_tree = np.array([[0, 1], [1, 2], [0, 3]])
populations_df = pd.DataFrame(populations, columns=["Id", "Step", "Pop"])
parent_tree_df = pd.DataFrame(parent_tree, columns=["ParentId", "ChildId"])
data = process_data(populations_df, parent_tree_df)
setup_figure()
fish_plot(*data)
plt.show()Calling the above code displays the following image:
Plots absolute population counts at each step.
| Base | --absolute |
|---|---|
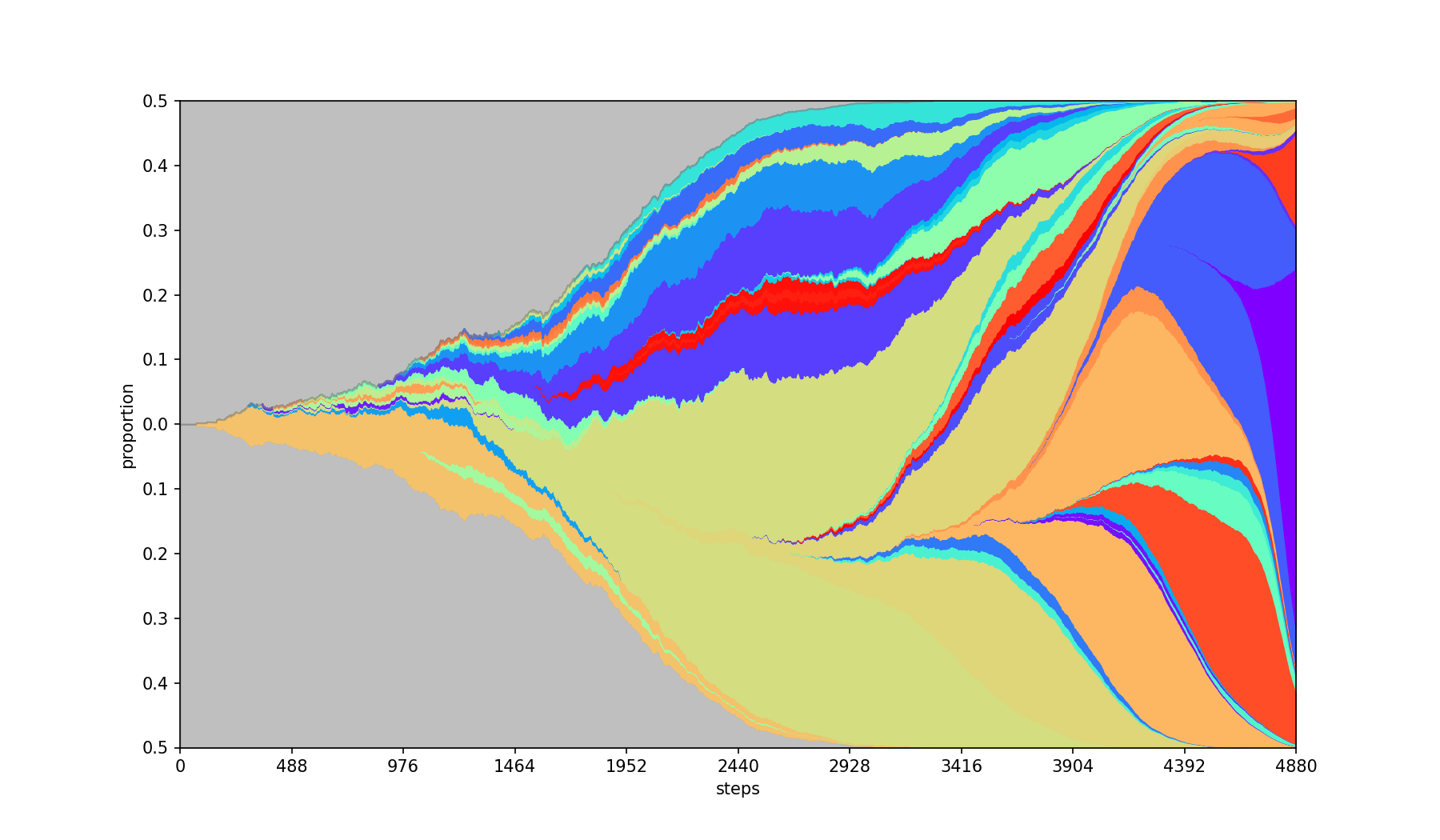 |
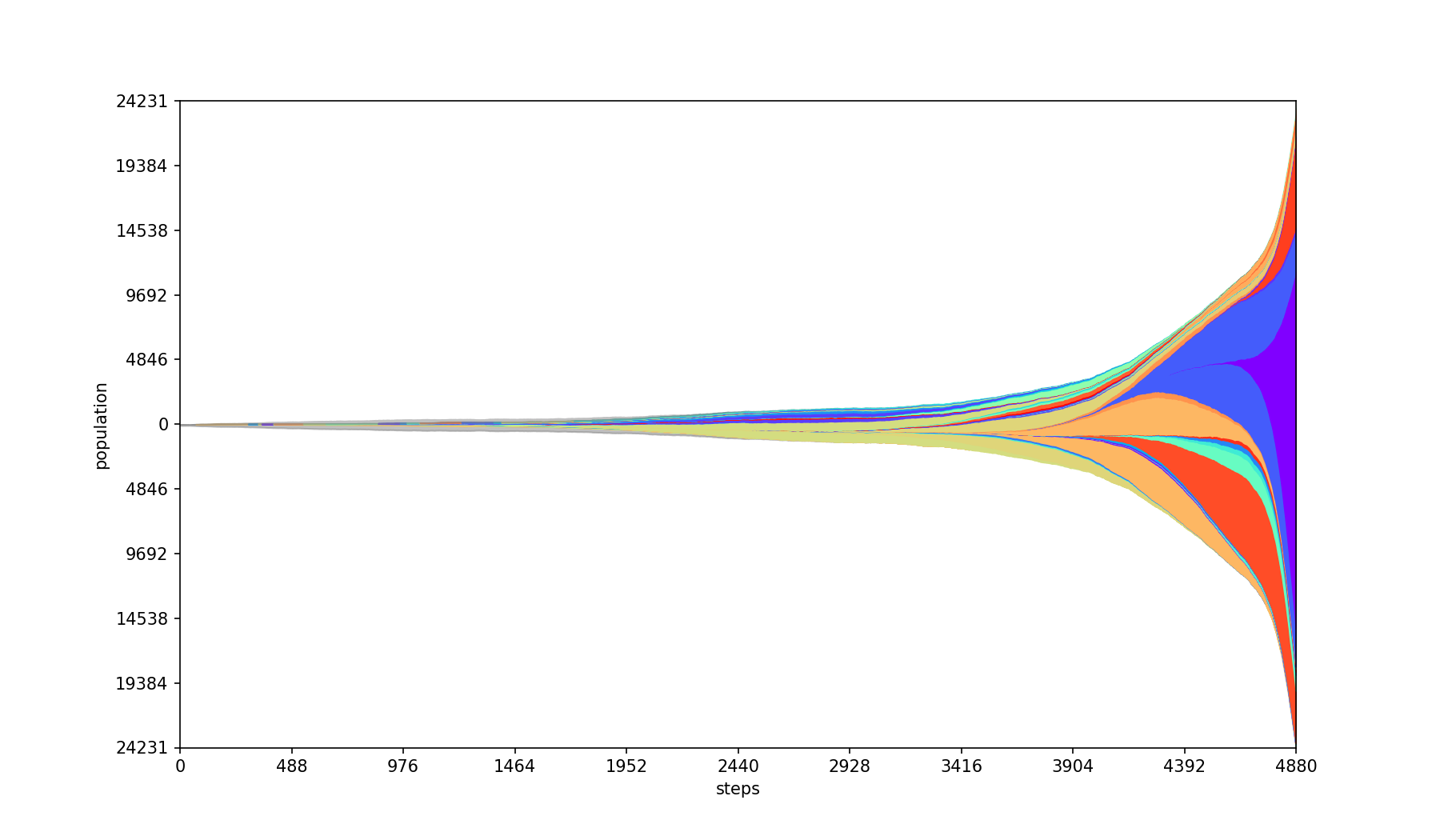 |
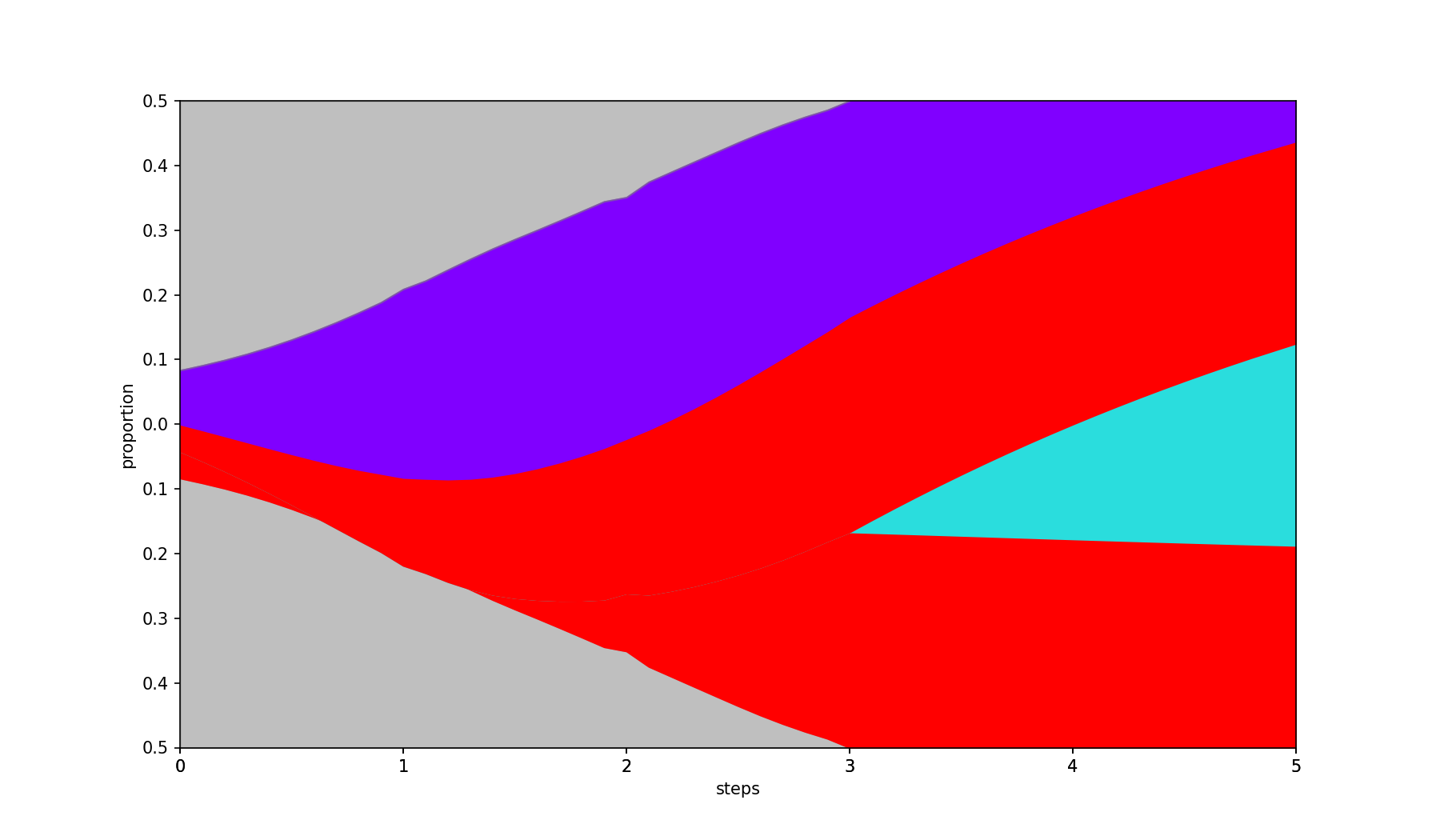
Fills in missing values by interpolation.
- A negative value (default) fills missing values with 0.
0uses linear interpolation between known data points.- A positive value uses polynomial (spline) interpolation of the given degree.
| Base | --interpolate 2 |
|---|---|
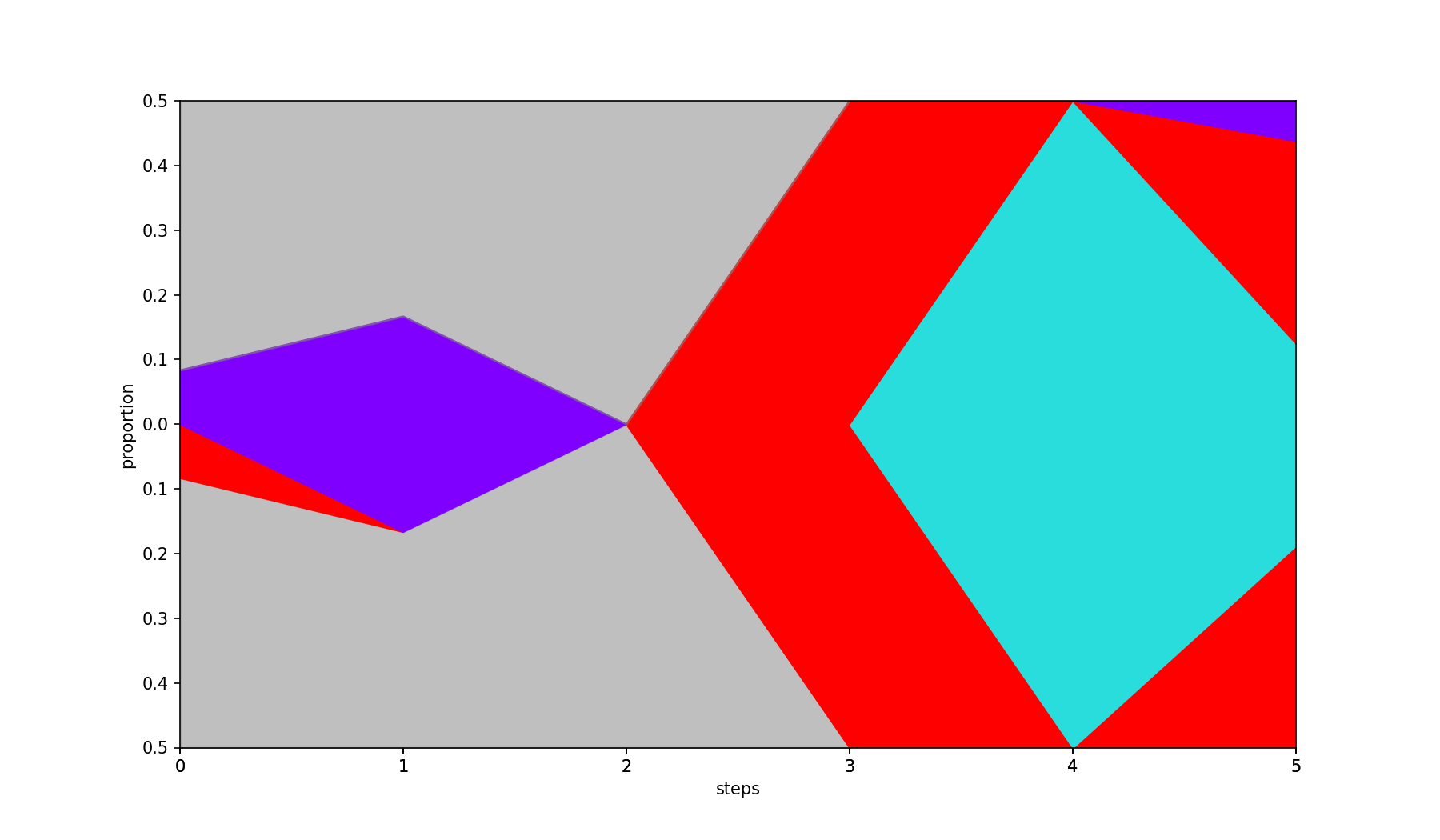 |
 |
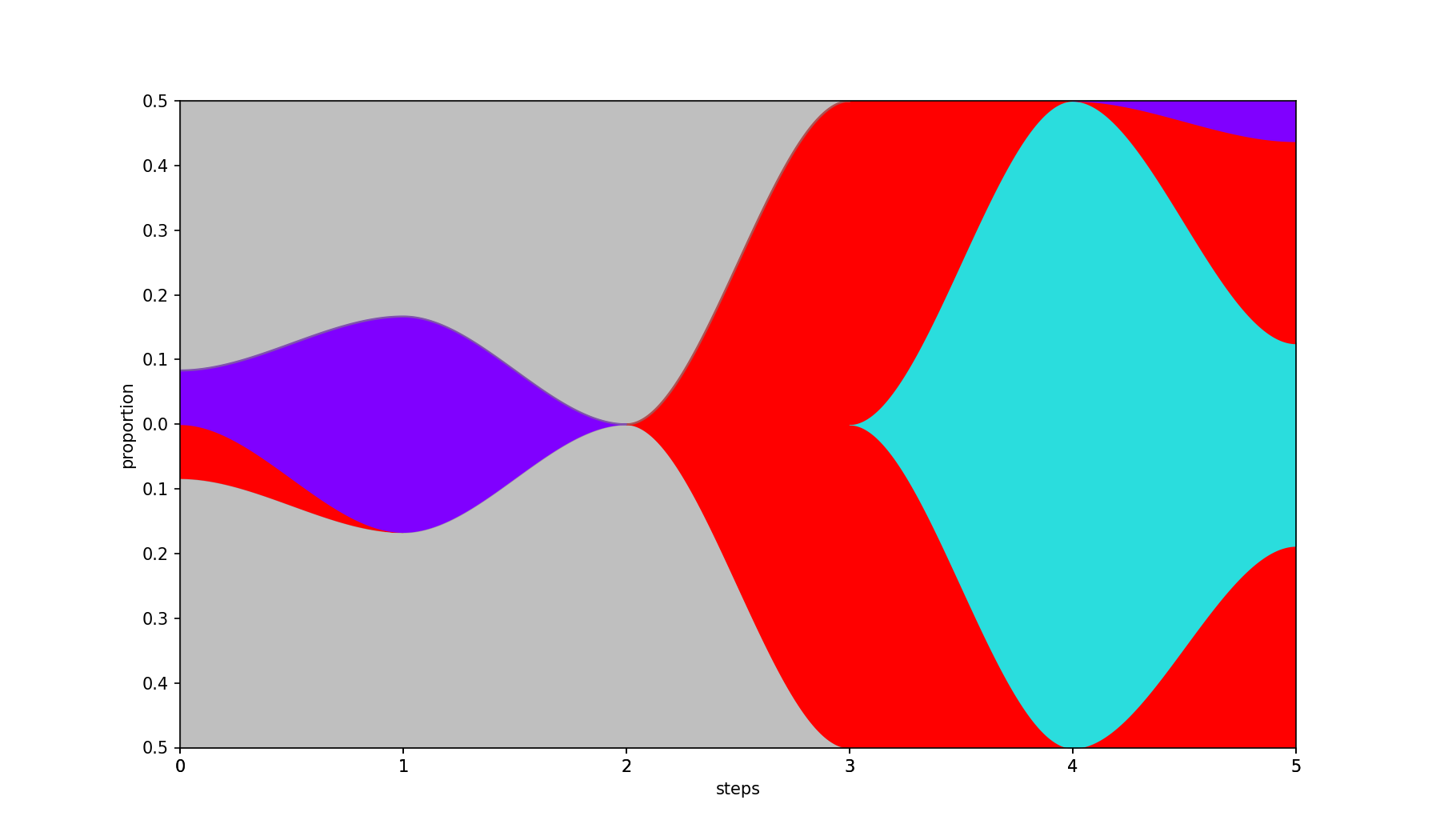
Smooths the filled areas using piecewise Hermite interpolation, producing gentle S-curve transitions between steps. Also adds a gray background and centers the plot when the population is empty at the first step.
| Base | --curved |
|---|---|
 |
 |
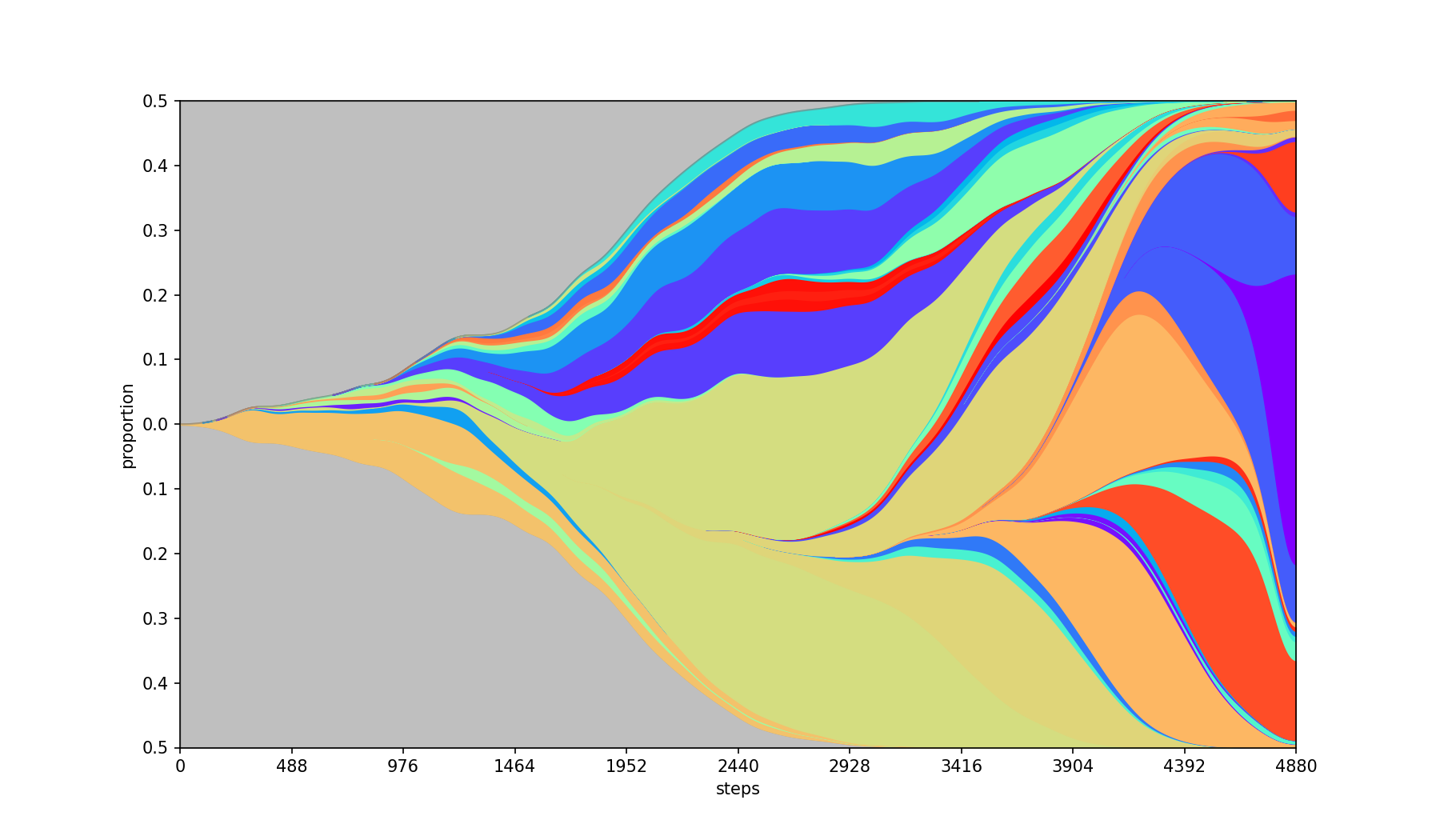
Smoothing of the graph using Gaussian filter. The parameter value is the standard deviation of the kernel. The bigger the population the bigger the value should be.
NOTE: If the population values are sparse, using smoothing without interpolation might lead to misleading population sizes.
| Base | --smooth 50 |
|---|---|
 |
 |
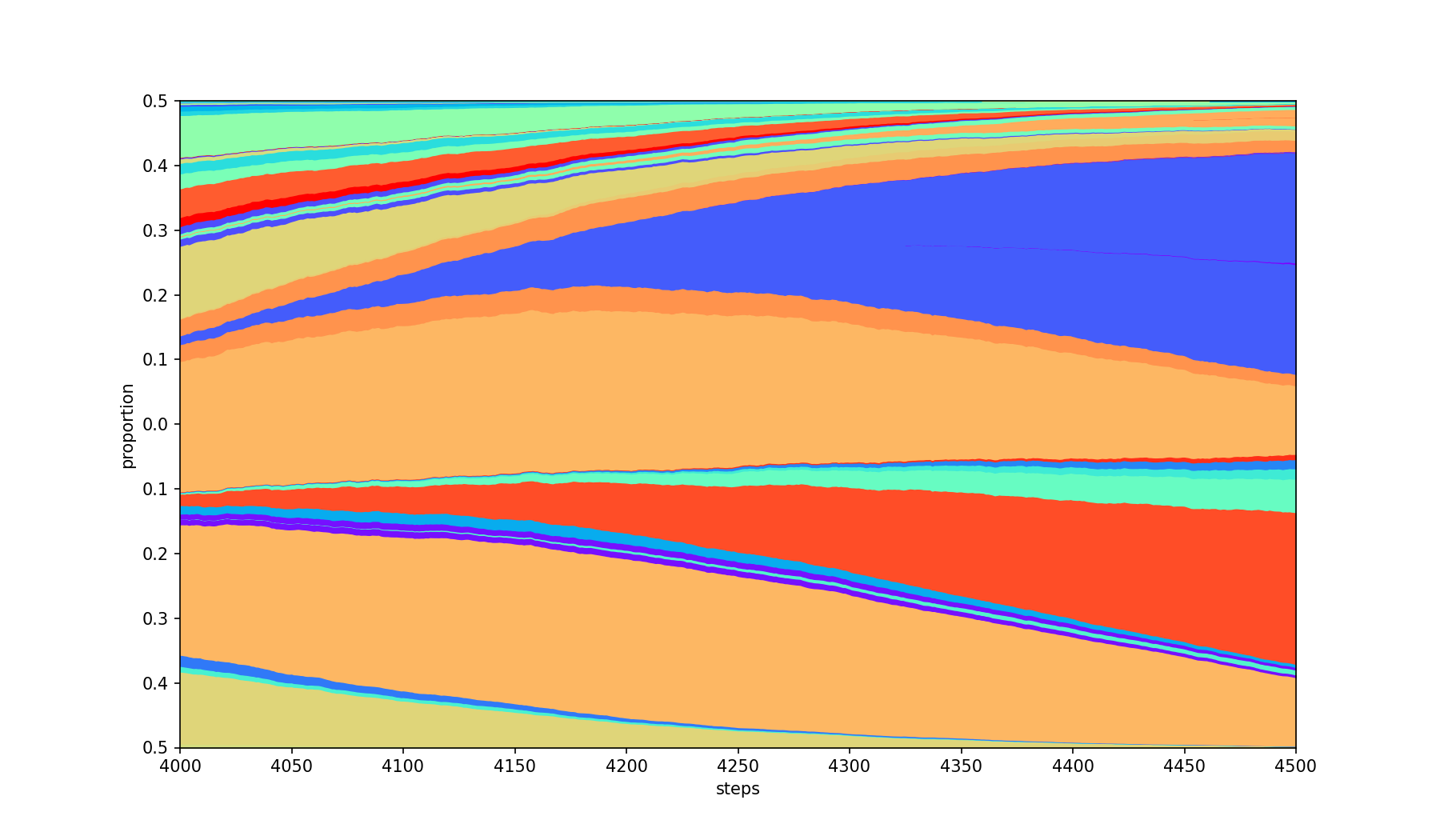
Only limits the steps to the range [first, last] inclusive.
| Base | --first 4000 --last 4500 |
|---|---|
 |
 |
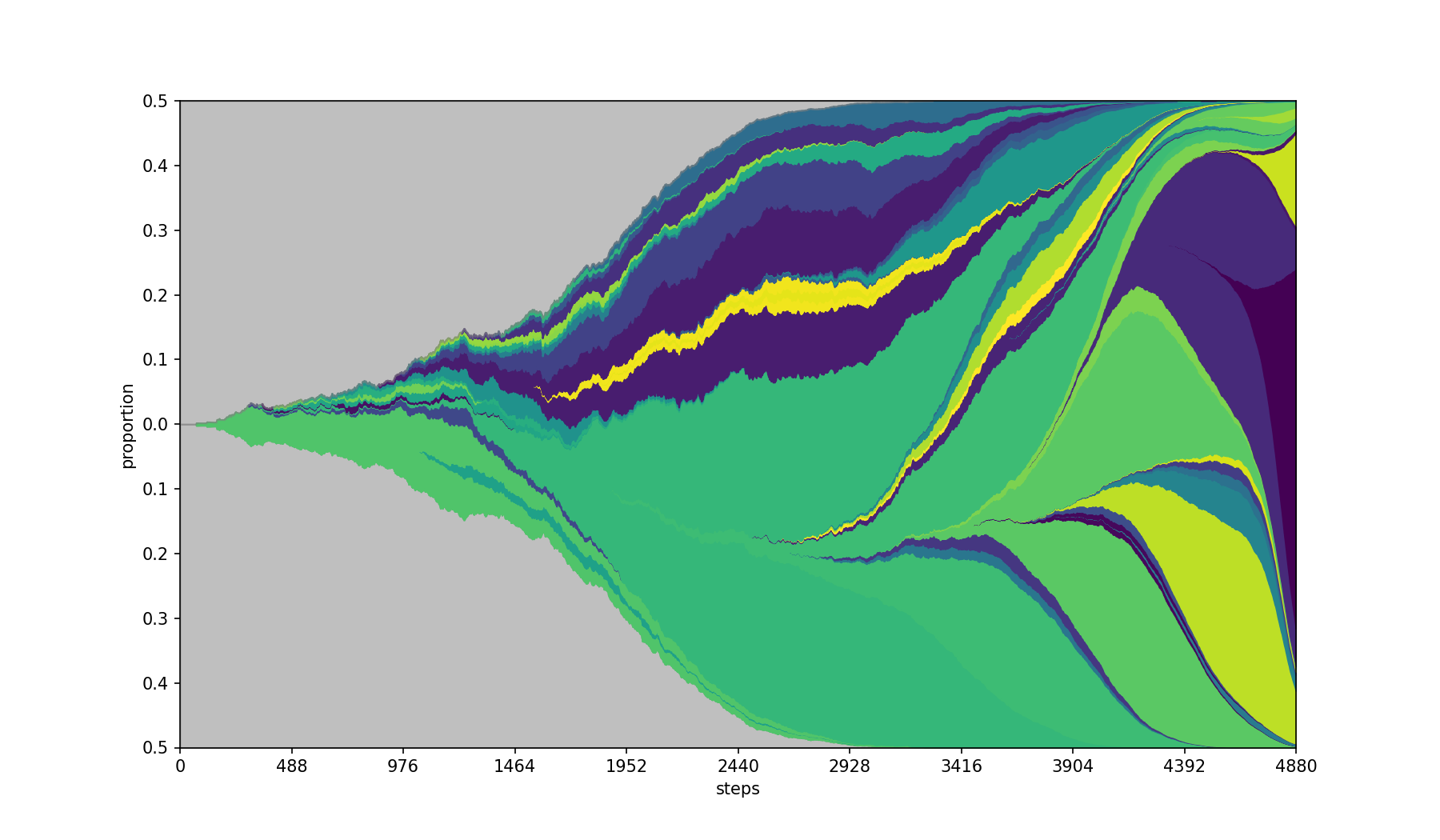
Use the specified matplotlib colormap.
Default colormap is rainbow.
| Base | --cmap viridis |
|---|---|
 |
 |
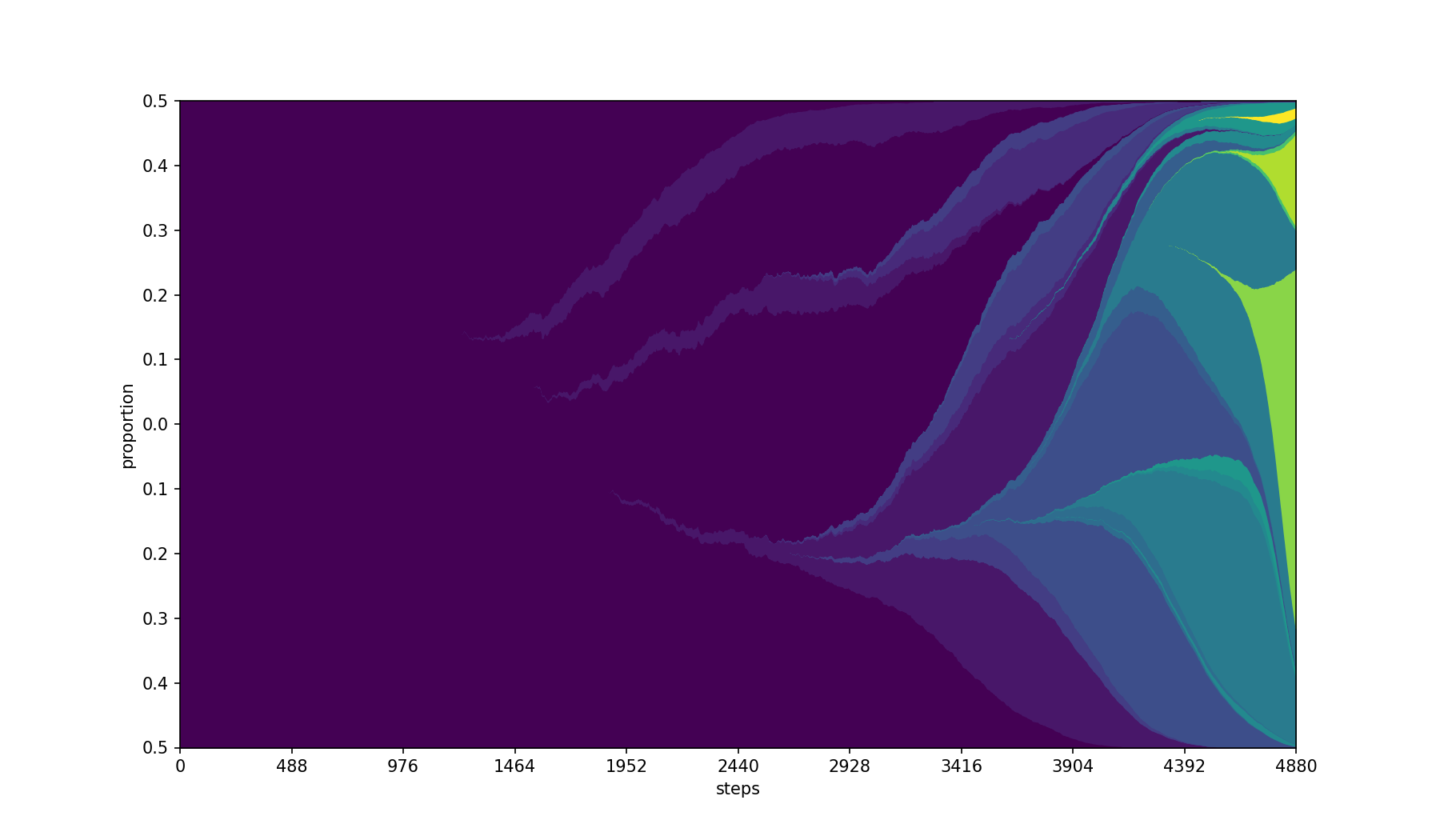
Color the ids based on a separate column in the populations.csv file. It will select the first value of the column per id, so the value should be constant for all entries with the same id.
Best combined with a sequential colormap using --cmap
| Base | --color-by Feature --cmap viridis |
|---|---|
 |
 |
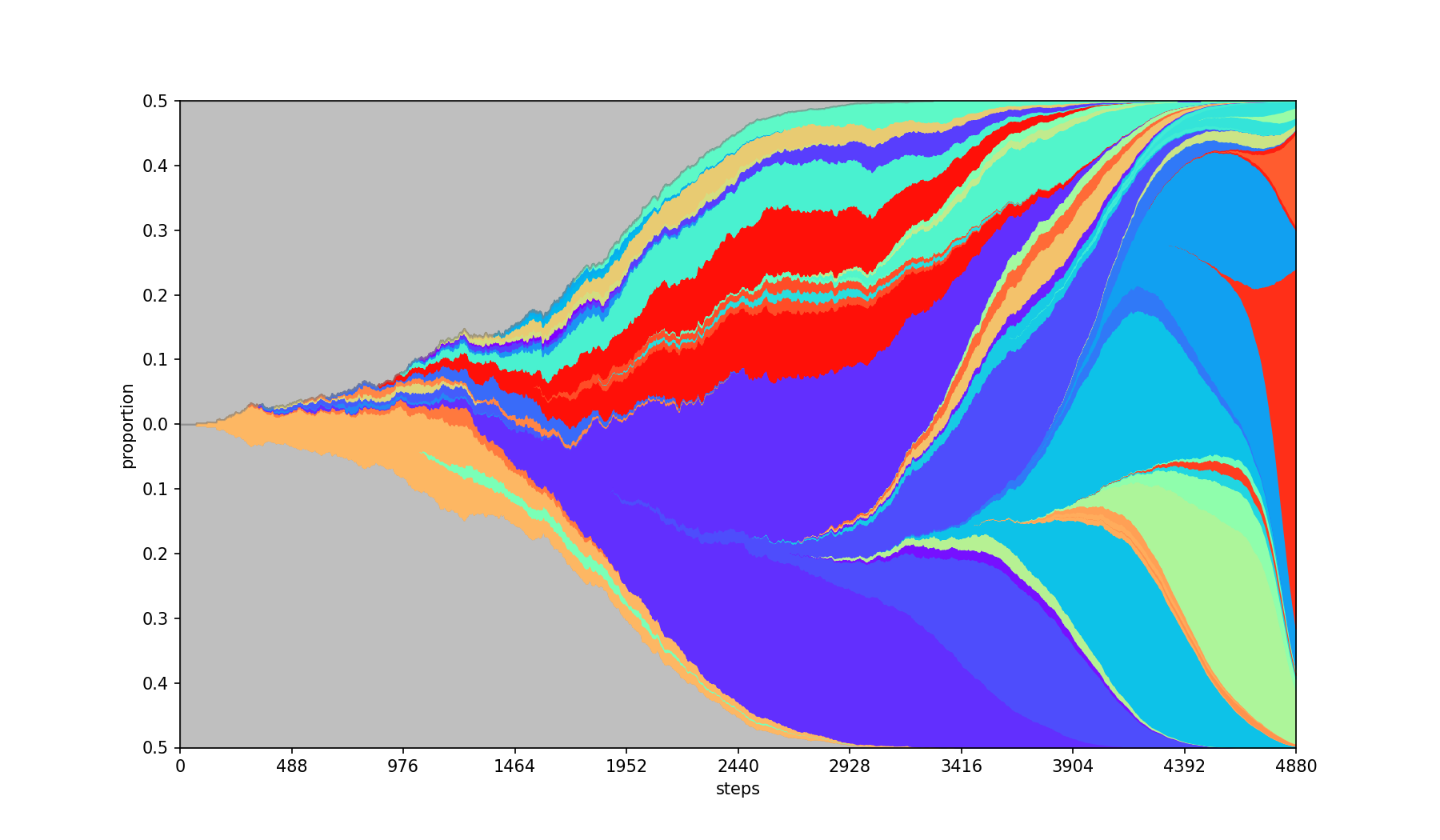
Specifies the seed for the randomization of colors.
| Base | --seed 2022 |
|---|---|
 |
 |
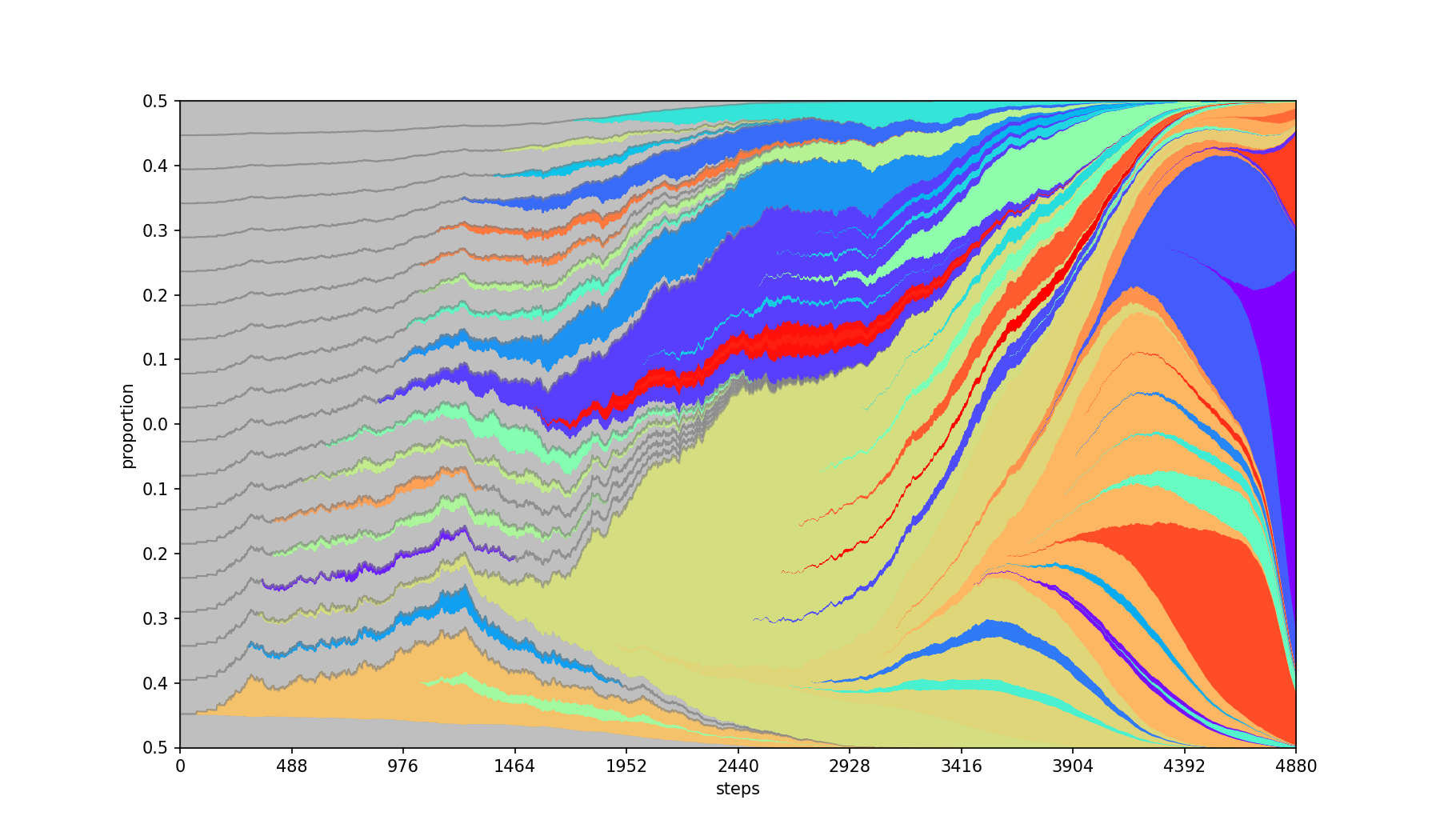
Places children equidistant from each other within their parent band. By default, children emerge from the center of the parent.
| Base | --separate |
|---|---|
 |
 |
Specifies the dimensions for the output image. The size is including the axes' labels.
Please cite as: Adam Streck, Tom L Kaufmann, Roland F Schwarz, SMITH: Spatially Constrained Stochastic Model for Simulation of Intra-Tumour Heterogeneity, Bioinformatics, 2023; https://doi.org/10.1093/bioinformatics/btad102
Authors: Adam Streck, Tom L. Kaufmann
Current contact: Adam Streck
Supervised by: Roland F. Schwarz
PyFish is available under the MIT License.
To actively develop the package, we recommend to install pyfish in development mode using pip pip install -e . --user.
In order to run the main routine from the command line without installing it first, run python -m pyfish.main -- tests/populations.csv tests/parent_tree.csv out.png.
To trigger testing, run pytest -v .
To generate the docs, run python ./tests/generate_doc_images.py